PiggyBac transposon vectors: the tools of the human gene encoding
Introduction
As the Nobel Prize winner, Renato Dulbecco, pointed out, the cellular genome must be concentrated on now if we wish to learn more about cancer. A genome contains an organism’s hereditary information. Concentrating on human genomemeans paying attentions to the hereditary information as well as the hereditary material. In order to uncover the code of life, the structure and function of the genes as well as the co-relationship among the genes need to be focused on from an overall level. Human genome project (HGP) was proposed by American scientists in 1985, and the project began in October 1990. The $3-billion project was founded in 1990 by the United States. In addition to the United States, the international consortium comprised geneticists in the United Kingdom, France, Australia, Japan and a myriad of other spontaneous relationships. The project was expected to take 15 years to identify all the human protein-coding genes, and to map the nucleotides contained in a human haploid reference genome (more than three billion). After the accomplishment of the HGP, the function of each gene was paid more attentions. The research strategy was to induce mutations in each single gene to observe their effects so that the molecular mechanism of gene function can be analyzed. However, because of the costly mammal-specific mutagens as well as larger required dose, traditional mutagen become costlier when applied to mammals, making it unavailable to large-scale analysis of gene function. Insertional mutagenesis using transposons makes genome-wide screens possible. The transposons can integrate with the genome and act as a tag, allowing its location to be determined and its effects on gene function to be assessed. However, transposon usage in mammals was limited due to the lack of efficient transposition system.
Mechanism of transposons
A DNA segment can be copied or cut down from its original position and insert into another site after cyclization to play a regulating role in its new position. This process is called transposition, and a DNA segment that undergoes transposition is a transposon. A transposon that inserts itself into a functional gene will most likely cause a mutation and dysfunction of that gene.
Transposons carry genes that code for transposition as well as for other functions. Transposons are assigned to one of two classes—retrotransposons and DNA transposons—according to different mechanism of transposition. Retrotransposons copy themselves first from DNA to RNA by transcription, then reverse transcript from RNA to DNA again, and the DNA copy is then inserted into the genome in a new position through a “copy-and paste” mechanism, while DNA transposons transpose via a “cut-and-paste” mechanism, without creating a second copy of the DNA during transposition (Figure 1). Since DNA transposons can be used as a kind of gene vehicles for the gene screening and gene function research, DNA transposon has become a research hotspot.
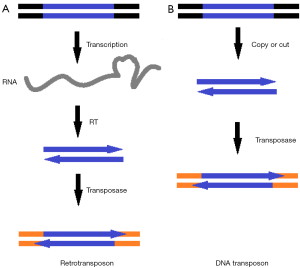
Transposons can be used to integrate transgenes into the host cell chromosome (1) or characterize specific gene function by disrupting gene function via insertion in or near genes (2). Besides, one of the features of transposons is that the transposase sequence can be separated from the transposon by genetic engineering. And the transposase can be transposition efficiency only when the transposon DNA fragment is available. Generating a binary transposition system with separated transposase can improve the security of transposition.
Advantages of piggyBac (PB) transposon
The usage of transposon in mammals has long been limited due to lack of efficient transposition system. Transposons that can be used in mammal cell include: (I) hAT-like Tol2, the only naturally active vertebrate transposon, isolated from the genome of the Japanese medaka fish; (II) Tcl-like transposons, Sleeping Beauty (SB) and Frog Prince, reconstructed from inactive transposons of fish and frog genomes, respectively; (III) PB isolated from the cabbage looper moth Trichoplusia ni. Among these transposon systems, SB and PB have the highest transposition activity in mammalian cells (3). In fact, two lines of evidence suggest that PB transposition may be a host factor-independent reaction: first, its transposition is highly efficient in a wide range of organisms such as yeast, insects, planarian, the malaria parasite, and mammals; second, transposition can be reconstituted in vitro by using purified piggyBac transposase and DNA elements (4,5). Compared with the PB transposon, the SB transposon often leave a CAG footprint for 3 base pair (bp), and has a tendency for local hopping. Its tendency for strong ‘local hopping’ and leaving footprints limit SB usage. PB has a stronger transposition activity than SB (6), and usually leaves no footprint after excision, which makes PB less likely to cause genomic damage during mutagenesis (7). This feature also makes PB transposon more popular. Since the discovery of the PB in cabbage looper moth Trichoplusia ni, it has mediated the transposition in Mediterranean fruit fly, Drosophila melanogaster, and silkworm, Bombyx mori. After that, people discovered that the PB transposon could also be used in mammal cells. Because of its high transposition activity and “footprint-free excision”, PB has created a new outlook for molecular medicine.
The PB system is derived from the cabbage looper moth Trichoplusia ni, and it’s an alternative transposon for gene delivery into mammalian cells. These transposable segments were initially discovered in mutant baculovirus strains, hence their name “PB”. The original PB element is approximately 2.4 kb with identical 13 bp terminal inverted repeats and additional asymmetric 19 bp internal repeats. A transgene can be inserted into The PB element, between the inverted repeat elements (8). And the PB transposase enzyme from a separate vector can insure the transposition activity. That is to say, the transposase and the sequence itself can be separated apart to build a binary transposition system with transposition activity. This arrangement permits a “cut-and-paste” transposition which can insert a transgene into the genome at TTAA segments (9). These features enable PB to be used for oncogene screening and genetic therapy.
Like the P-element, the PB transposon is also a kind of transposon that transpose through a “cut-and-paste” mechanism. However, unlike the P-element, PB can precisely complete excision and reinsertion in many insect chromosomes; both the excision and the reinsertion require the transposase coding by PB itself, and occurs in the TTAA target sites (10). The PB element can be cut down from the donor chromosome by the transposase, and the split donor DNA was then reconnected with DNA ligase. After that, the vector was inserted into the target insertion site (TTAA), complete the transposition (Figure 2). It is unique that no DNA synthesis occurs during the PB transposition, and the target gaps caused by transposition are sealed simply by ligation, making the transposition of PB precise and defining (11).
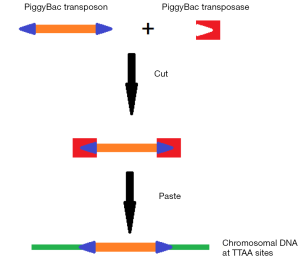
PB has many priorities to other current vectors. Compared with virus vectors, PB has a higher security and is more convenient to generate. Compared with other transposons, PB has a larger cargo (approximately 9.1–14.3 kb), a higher transposition activity, and its footprint-free characteristic makes it more practical. As a new transgene vector, PB has these priorities: (I) high efficiency transposition; (II) large cargo; (III) steady long-term expression; (IV) the trans-gene is integrated as a single copy; (V) tracking the target gene in vivo by a noninvasive mark instead of traditional method such as PCR; (VI) easy to determine the integration site.
As a mutagen, PB also has these priorities: (I) PB transposon is able to transposition in reproductive cells effectively, which means that it is possible to generate a single PB inserted animal strain efficiently, instead of using stem cell culturing, surgery, and other traditional mutagenesis methods; (II) PB insertional site can be easily located by inverse PCR; (III) the PB insertion is widely distributed in the genome, with a preference to gene regions; (IV) PB insertion can efficiently disable the endogenous gene, performing a similar phenotype to traditional mutagenesis; (V) precise excision of PB enable recovery of mutation to confirm the association between genotype and phenotype; (VI) it is possible to introduce visible genetic marker to track the transposon and transposase so that distinguishing homozygous from heterozygous is possible by naked eyes; (VII) combined with gene trapping to screen the expression profiling of the endogenous gene. Thus, PB transposon is not only a transgene tool, but also a potential non-virus vector, which can be widely used in scientific experiments.
PB as a gene vector
Currently used transgene vectors include virus, transposon etc. PB transposon has several priorities to other current vectors. It has higher security than virus vectors, and it’s more convenient to be generated (12). PB can also move larger DNA fragment (9.1–14.3 kb) than other transposons, so PB has a higher transposition activity (13,14).
PB has a 13 bp perfect terminal invert repeat sequence. PB can be inserted into TTAA segments, and can also be excised in mammals precisely, which makes PB a transgene vector with great expectation (15-17).
Besides, among all the transposition systems, PB is the transposon most suitable for chimeric transposition. Claudia Kettlunand his crew fused a highly site-specific synthetic zinc-finger protein (ZFP) DNA-binding domain to the N-terminus of the PB transposase, and evaluated site-directed genomic integration in human cells (18). By inserting a DNA binding structure into the PB transposon, PB can be used to mediate site-directed genomic integration. PB has also been used to generate human and mouse induced pluripotent stem cells (iPS cells) (19-22). These features make PB a potential tool for future trans genetic experiment.
PB for mammal gene function
There are approximately 28,000 protein coding genes in the human genome, excluding many non-coding RNA and DNA regulation elements. It is anticipated that further information of the human genome will provide new avenues for advances in medicine and biotechnology. Genetic analyzing on mammals would reveal the function of genes and discover pathogenic genes. Sheng D and his staff have generated an efficient PB transposition system in mammalian cells and mice (23). Instead of stem cell culturing, this technique directly reforms the genetic material in fertilized egg. It took the researchers 3 months to determine the function of 70 mouse genes. Besides the high efficiency of PB in mammals, the mutation of PB can also be reported and tracked by luciferase, enable the researchers to identify the PB insertional mutation easily (24). Researchers have also established a data base storing the PB insertions in the mouse genome (25). The first high efficiency mammal TEs system was built in 2005, providing an efficient genome level mutagen tool for large scale research on mammal gene function, creating a new outlook for human gene therapy.
PB for screening cancer genes
Because of the lack of efficient insertional mutagenesis tools, gene screening in higher organisms has been hampered for decades. Retroviruses have been used for cancer gene discovery in mice, but their application has been limited to the study of hematopoietic and mammary tumors because of viral tropism for these tissues. After that, people discovered that SB is able to transpose in multiple cell types, including mouse embryonic stem cells. Another transposon, PB, has been shown to have a weaker tendency for local hopping in vitro. PB can also move larger DNA fragments than SB. Furthermore, in contrast to SB, PB does not leave undesired footprint mutations after transposition. In summary, PB is a better tool for genetic screening.
Inserting potential oncogene into the genome through PB can efficiently induce cancer developing. Roland Rad and colleagues generated PB transposase knockin mice and mouse lines carrying transposons (2). They found that, beginning at 2 months of age, double-transgenic mice developed cancers, the results demonstrate the unique qualities of PB for genome-wide mutagenesis. Apart from oncogene insertions, hits in tumor suppressor genes can also be used to reveal the function in the disease-causing mechanism of the targeted gene. The activation of the oncogene or the inactivation of the tumor suppress gene caused by PB insertion in cancer genes will both induce the tumor growth. The location of cancer genes can then be identified by splinkerette PCR or other methods that can map the PB insertional sites (26).
PB for genetic therapy
Efficient tools are necessary for gene therapy to be put into clinical use, by providing the stable delivery of genetic information into eukaryotic genomes. Most current gene delivery strategies are based on viral vectors. However, various of hinders, such as the limited cargo capacity, host immune response and mutational risks, highlight the need for alternative gene delivery tools (27).
The extraordinary performance of PB transposon gives us a new choice. PB has a higher transposition activity than SB in human cell lines, while overproduction inhibition was not observed with PB, while it is a major limitation of the SB system. Hideyuki Nakanishi and his staff succeeded in Transposition in human hepatocyte-derived cell lines with PB, achieved a prolonged expression (28). PB is the most promising transposon-based vector system for achieving site-specific targeting of therapeutic genes due to the flexibility of its transposase for being molecularly engineered (29,30).
Conclusions
As the rapid development of transgene technology, people have a higher requirement for the gene vector. The focus of molecular medicine researcher has moved to a more efficient way for gene screen and gene therapy with transposons. Due to its high cargo capacity and security, PB has become an optional tool for mammal transposition. As the cognition of its function, the PB transposon will be applied in more areas such as gene function and therapy, transposon mutagenesis of cell line, engineered protein expression and so on.
Acknowledgements
None.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
References
- Wilson MH, Coates CJ, George AL Jr. PiggyBac transposon-mediated gene transfer in human cells. Mol Ther 2007;15:139-45. [PubMed]
- Rad R, Rad L, Wang W, et al. PiggyBac transposon mutagenesis: a tool for cancer gene discovery in mice. Science 2010;330:1104-7. [PubMed]
- Kim A, Pyykko I. Size matters: versatile use of PiggyBac transposons as a genetic manipulation tool. Mol Cell Biochem 2011;354:301-9. [PubMed]
- Yusa K, Zhou L, Li MA, et al. A hyperactive piggyBac transposase for mammalian applications. Proc Natl Acad Sci U S A 2011;108:1531-6. [PubMed]
- Burnight ER, Staber JM, Korsakov P, et al. A Hyperactive Transposase Promotes Persistent Gene Transfer of a piggyBac DNA Transposon. Mol Ther Nucleic Acids 2012;1:e50. [PubMed]
- Wu SC, Meir YJ, Coates CJ, et al. piggyBac is a flexible and highly active transposon as compared to sleeping beauty, Tol2, and Mos1 in mammalian cells. Proc Natl Acad Sci U S A 2006;103:15008-13. [PubMed]
- Wang W, Lin C, Lu D, et al. Chromosomal transposition of PiggyBac in mouse embryonic stem cells. Proc Natl Acad Sci U S A 2008;105:9290-5. [PubMed]
- Cadiñanos J, Bradley A. Generation of an inducible and optimized piggyBac transposon system. Nucleic Acids Res 2007;35:e87. [PubMed]
- Maragathavally KJ, Kaminski JM, Coates CJ. Chimeric Mos1 and piggyBac transposases result in site-directed integration. FASEB J 2006;20:1880-2. [PubMed]
- Keith JH, Schaeper CA, Fraser TS, et al. Mutational analysis of highly conserved aspartate residues essential to the catalytic core of the piggyBac transposase. BMC Mol Biol 2008;9:73. [PubMed]
- Mitra R, Fain-Thornton J, Craig NL. piggyBac can bypass DNA synthesis during cut and paste transposition. EMBO J 2008;27:1097-109. [PubMed]
- Park TS, Han JY. piggyBac transposition into primordial germ cells is an efficient tool for transgenesis in chickens. Proc Natl Acad Sci U S A 2012;109:9337-41. [PubMed]
- Wilber A, Linehan JL, Tian X, et al. Efficient and stable transgene expression in human embryonic stem cells using transposon-mediated gene transfer. Stem Cells 2007;25:2919-27. [PubMed]
- Li R, Zhuang Y, Han M, et al. piggyBac as a high-capacity transgenesis and gene-therapy vector in human cells and mice. Dis Model Mech 2013;6:828-33. [PubMed]
- Liu X, Li N, Hu X, et al. Efficient production of transgenic chickens based on piggyBac. Transgenic Res 2013;22:417-23. [PubMed]
- Elick TA, Bauser CA, Fraser MJ. Excision of the piggyBac transposable element in vitro is a precise event that is enhanced by the expression of its encoded transposase. Genetica 1996;98:33-41. [PubMed]
- Lee CY, Li JF, Liou JS, et al. A gene delivery system for human cells mediated by both a cell-penetrating peptide and a piggyBac transposase. Biomaterials 2011;32:6264-76. [PubMed]
- Kettlun C, Galvan DL, George AL Jr, et al. Manipulating piggyBac transposon chromosomal integration site selection in human cells. Mol Ther 2011;19:1636-44. [PubMed]
- Kim D, Kim CH, Moon JI, et al. Generation of human induced pluripotent stem cells by direct delivery of reprogramming proteins. Cell Stem Cell 2009;4:472-6. [PubMed]
- Yusa K, Rad R, Takeda J, et al. Generation of transgene-free induced pluripotent mouse stem cells by the piggyBac transposon. Nat Methods 2009;6:363-9. [PubMed]
- Woltjen K, Michael IP, Mohseni P, et al. piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 2009;458:766-70. [PubMed]
- Tsukiyama T, Asano R, Kawaguchi T, et al. Simple and efficient method for generation of induced pluripotent stem cells using piggyBac transposition of doxycycline-inducible factors and an EOS reporter system. Genes Cells 2011;16:815-25. [PubMed]
- Ding S, Wu X, Li G, et al. Efficient transposition of the piggyBac (PB) transposon in mammalian cells and mice. Cell 2005;122:473-83. [PubMed]
- Landrette SF, Cornett JC, Ni TK, et al. piggyBac transposon somatic mutagenesis with an activated reporter and tracker (PB-SMART) for genetic screens in mice. PLoS One 2011;6:e26650. [PubMed]
- Sun LV, Jin K, Liu Y, et al. PBmice: an integrated database system of piggyBac (PB) insertional mutations and their characterizations in mice. Nucleic Acids Res 2008;36:D729-34. [PubMed]
- Chew SK, Rad R, Futreal PA, et al. Genetic screens using the piggyBac transposon. Methods 2011;53:366-71. [PubMed]
- Claeys Bouuaert C, Chalmers RM. Gene therapy vectors: the prospects and potentials of the cut-and-paste transposons. Genetica 2010;138:473-84. [PubMed]
- Nakanishi H, Higuchi Y, Kawakami S, et al. piggyBac transposon-mediated long-term gene expression in mice. Mol Ther 2010;18:707-14. [PubMed]
- Meir YJ, Weirauch MT, Yang HS, et al. Genome-wide target profiling of piggyBac and Tol2 in HEK 293: pros and cons for gene discovery and gene therapy. BMC Biotechnol 2011;11:28. [PubMed]
- Feschotte C. The piggyBac transposon holds promise for human gene therapy. Proc Natl Acad Sci U S A 2006;103:14981-2. [PubMed]


