The role of SOX2 in small cell lung cancer, lung adenocarcinoma and squamous cell carcinoma of the lung
Sex-determining region Y (Sry)-related high mobility group (HMG)-box (SOX) genes are indispensable for multiple aspects of development (1). Members of the SOX family are expressed in a wide variety of tissues and have important roles in the regulation of organ development and cell-type specification (1). SOX2 was initially reported to be strongly associated with the inhibition of neuronal differentiation (2). More recent studies indicate that SOX2 exists in the nuclei of embryonic stem (ES) cells and acts as a transcriptional factor to maintain their unique characters such as clonogenicity, pluripotency, and self-renewal (3). Some cancer stem cells (CSCs) have, on their cell surface, ATP-binding cassette transporters (ABCG) that pump out the DNA-binding dye Hoechst 33342 and are characterized as side population (SP) cells (4). Because these malignant SP cells proliferate in a sustained fashion and readily export many cytotoxic drugs, they may be resistant to therapy and contribute to disease relapse (5). It was found that isolated SP cells show higher expression levels of stem cell genes, such as SOX2 and Oct4 and tumorigenesis properties than non SP cells (6).
SOX2 is transcriptionally regulated by an enhancer containing a composite SOX-OCT element that the octamer-binding transcription factor 4 (Oct4) and SOX2 bind in a combinatorial interaction (7). It appears that the SOX2-Oct4 regulatory complex upregulates a large number of genes important for the maintenance of the pluripotency of ES cells and downregulates genes responsible for the initiation of differentiation (8). There is abundant evidence that SOX proteins might also affect the Wnt/β-catenin pathway. They can either antagonize or facilitate β-catenin/TCF-mediated transcription in the context of different SOX species (9). In parallel, G1/S-specific cyclin-D1 (CCND1) has been identified as the downstream target of SOX2 which agrees well with the cellular behavior of SOX2 in promoting the G0/G1 to S transition (10). Above all, SOX2 has been recently recognized as a novel target of EGFR-Src-AKT signaling in NSCLCs that modulates self-renewal and expansion of stem-like cells, making the relative SOX2 expression and functions within the tumor-CSCs a major determinant in EGFR-targeted therapy (11).
A number of links have been found between SOX transcription factors and human cancers. For instance, SOX2 promoter silencing by DNA methylation has been reported in some human gastric carcinomas (12). In contrast, several publications report overexpression of SOX2 in glioblastomas (13), non-small cell lung cancer (NSCLC) (14,15), SCLC (16), prostate cancer (17), hepatocellular carcinomas (18) osteosarcomas (19), and breast carcinomas (20), supporting a role of SOX2 as an oncogene in these tissues. These reports suggest that SOX2 could activate important gene cascades involved in tumor initiation and progression and in the maintenance of a poorly differentiated state. In this review, we will attempt to deepen our knowledge on the underlying molecular mechanism of SOX2’s function in lung tumorigenesis, which may emerge as a novel promising strategy for lung cancer therapy.
The role of SOX2 in small cell lung cancer
SCLC is a distinct clinical and histological entity within the range of lung cancer (21). The incidence and mortality of SCLC worldwide make this disease a notable health-care issue. SCLC represents 13% of all newly diagnosed cases of lung cancer worldwide, and its prognosis remains poor, with an overall median survival following treatment of 10 months and a 5-year survival of 5% (22). Its management has followed the major developments of modern cancer treatment through the integration of biology, imaging, chemotherapy, and radiotherapy.
The immunohistochemical analysis of SOX2 expression in various types of lung cancer found that SCLC tissues revealed a higher expression level of SOX2 than NSCLC tissues (23). In parallel, SOX2 was found to cooperate with important oncogenes like Wnt1, Wnt2, c-Myc and Notch to promote lung tumor occurrence, while downregulation of SOX2 inhibited proliferation and induced apoptosis in tumor cells (23). Another study of more than 50 tumour samples and SCLC cell lines (H446 and H720) has shown that SOX2 is amplified in approximately 27% of cancers (16).
SOX2 plays a pivotal role in the maintenance of ES cell pluripotency by regulating lineage commitment factors and later in development, is involved in specification and maintenance of neural stem cells during neurogenesis. Notably, conditional induction of SOX2 in lung epithelial cells is also known to increase the number of neural progenitor cells (24). SCLCs are tumors with neuroendocrine features. SOX2 protein overexpression has previously been noted in high-grade SCLC, and immunoreactive antibodies against SOX2 have been detected in sera from SCLC patients (25,26).
Two possibilities may account for the increased expression of SOX2 in SCLC. One is that the normal progenitor cell of SCLC, generally presumed to be the neuroendocrine Kulchitsky cell, expresses SOX genes; thus, the expression of this antigen in SCLC represents the persistence of these differentiation characteristics during neoplastic clonal expansion. The other possibility is that SOX genes are not expressed in normal adult Kulchitsky cells or bronchial epithelium and that the expression of these genes in SCLC represents a reactivation of lineage-specific embryonic markers, reflecting the developmental stage at which SOX2 is coexpressed. Rudin et al., found that cell proliferation can be suppressed in vitro by silencing SOX2 (using short hairpin RNAs) which implicates this gene in driving SCLC and suggests a plausible novel therapeutic strategy (16).
The role of SOX2 in squamous cell lung cancer
It has been found that SOX2 amplification and consequent SOX2 protein overexpression represent important mechanisms of tumor initiation and progression in a considerable subset of squamous cell carcinomas (SCCs) (27). The reported frequency of SOX2 amplification in lung SCCs varies from 20% to 60% with these variations in frequency being most likely due to methodological discrepancies applied by different laboratories but also differences between the cohorts and tumor heterogeneity (28-31). Lung SCCs are known to commonly harbor an amplification of the genomic region 3q. The amplification locus of 3q comprises additional genes, such as the defective in cullin neddylation 1, domain containing 1 (DCUN1D1) and the phosphoinositide-3-kinase, catalytic, alpha polypeptide (PIK3CA) that previously have been proposed to be target oncogenes of the 3q amplicon. Recent studies have provided strong evidence that SOX2 is the primary amplification target within the common 3q amplicon and functional studies using shRNA against SOX2 showed an impact on tumor biology, thus making SOX2 the most promising candidate 3q oncogene (28,29,32).
A peak of genomic amplification on chromosome 3q26.33 that contains SOX2 gene, in SCCs of the lung and esophagus has been recently reported (28). However SOX2 alone cannot transform immortalized tracheobronchial epithelial cells and Forkhead box E1(FOXE1) or Fibroblast growth factor receptor 2 (FGFR2) are required as transforming cooperative genes (28). Toschi et al., evaluated SOX2 and FGFR1 gene copy number by fluorescence in situ hybridization (FISH) in tissue microarray cores in 447 surgically resected NSCLCs, to investigate their prognostic relevance and their association with clinico-pathological characteristics. They reported that increased SOX2 and FGFR1 gene copy number is a common event in lung cancer patients with squamous cell histology and that SOX2 gene gain is a favorable prognostic factor in early stage resected patients (33).
It seems that SOX2 correlates with markers of squamous differentiation in lung SCCs. For instance, TP63 and Keratin 6A (KRT6A), which encode for the squamous markers p63 and cytokeratin 6A, respectively, were among the transcripts most correlated with SOX2 expression in lung SCCs (28). When SOX2 was ectopically expressed in the NCI-H2009 lung adenocarcinoma cell line, both TP63 and KRT6A were induced, demonstrating actions of SOX2 that promote squamous identity rather than de-differentiation to a pluripotent state, thus consistent with its role as a lineage survival oncogene (28). It has also been demonstrated that SOX2 overexpression in epithelial cells of the adult lung drives development of histologically well differentiated adenocarcinoma with significant squamous cell features including widespread expression of p63, FoxE1 and Desmoglein-3, a phenotype not unique to tumors induced by SOX2 (14). Inducible deletion of the tumor suppressor liver kinase B1 (LKB1) along with inducible expression of oncogenic K-Ras leads to adenocarcinoma with squamous features as well (34). It is possible that SOX2 alone can drive expression of some squamous tumor markers, but an additional oncogenic stimulus is required to drive complete squamous differentiation.
Within the primitive foregut there is reciprocal expression of NKX2.1 (also known as thyroid transcription factor 1; TITF1) and SOX2 in compartments that form the trachea and esophagus, respectively (35). As the developmental transcription factor NKX2-1 is an amplified lineage survival oncogene in lung adenocarcinoma, SOX2 may similarly represent a lineage survival oncogene in lung SCCs (35). Bass et al., found that SOX2 amplification was enriched in the lung SCC tumor population, while NKX2-1 amplification was enriched in lung adenocarcinoma (28). The complementary roles of SOX2 and NKX2-1 in distinct cancer lineages thus parallel their actions in development.
Functional studies underscore the oncogenic role of SOX2 in squamous cell carcinomas. In squamous cell carcinoma cell lines harboring SOX2 amplification, suppression of SOX2 had an anti-proliferative effect (28). Furthermore, cell lines overexpressing SOX2 exhibited increased migratory activity and enhanced colony formation (29). In preinvasive lesions of the lung, SOX2 expression has been reported to occur in normal bronchial epithelium, alveolar bronchiolization, squamous dysplasia, as well as carcinoma in situ (30). Furthermore, SOX2 amplification was reported in none of a series of low-grade bronchial lesions, but in all high-grade lesions, suggesting upregulation during preinvasive disease progression (36). Consistently, it has been shown that conditional homozygous SOX2 overexpression in Clara cells induces bronchial epithelial hyperplasia with 50% of cases showing a progression to lung cancer in mice (14). Taken together, these results strongly indicate that SOX2 harbors oncogenic potential and has a role during tumorigenesis.
An association between elevated SOX2 expression and indicators of better patient outcome, most importantly prolonged overall survival, was recently demonstrated (37). Increased levels of SOX2 amplification indicated a better histological differentiation grade and a trend to improved patient survival (37). In a cohort of early stage lung SCCs, patients with SOX2 expression above the median showed prolonged overall survival (14). The molecular mechanisms accounting for SOX2 being associated with favorable prognosis in lung SCCs is still unknown and further studies are needed to clarify the functional aspects of SOX2. SOX2 overexpression might recapitulate transcription networks active in normal squamous precursor cells and thus counteract the chaos of malignant dedifferentiation or alternatively, SOX2 overexpression might occur early during lung SCC carcinogenesis and might be lost during disease progression, due to genetic inactivation. Furthermore tumors arising from an upregulation of SOX2 exhibit a clear squamous cell differentiation and thus can be associated with better prognostic features, similar to NKX2-1 in lung adenocarcinomas.
Besides the lung, SOX2 has been found to be amplified and expressed in squamous cell carcinomas originating from other organ sites, predominantly derived from the embryonic foregut, but also from non-foregut tissues, such as the skin, the cervix, and the penis (27,28,38,39). Squamous carcinogenesis from diverse body sites may thus share similar underlying mechanisms and SOX2 might be a general marker for SCC differentiation regardless the tissue of origin. All the above studies used a similar strategy of chromosomal aberrations screening to identify the SOX2 locus as one of the most frequently amplified sites over the SCC genome and further highlighted the recurrent SOX2 activation and its indispensable role for squamous cell survival. However, it remains to assess the impact of the recurrent activation of SOX2 in advanced primary tumors and how SOX2 may mechanistically be involved in tumor progression and aggressiveness.
The role of SOX2 in lung adenocarcinomas
Previous findings revealed that SOX2 is expressed in bronchial epithelial cells of the lung, whereas it is absent in alveolar cells (30,38). Likewise, adenocarcinoma precursor lesions, such as atypical adenomatous hyperplasia, proved to be negative for SOX2 expression (30). In the study of Cai et al., the amplification of SOX2 in SCCs and adenocarcinomas was 31.6% and 20%, respectively (40). No SOX2 amplification was found among smokers with adenocarcinoma (40). In contrast, 10 of 38 (26.3%) cases involving patients with no history of smoking and with adenocarcinoma presented SOX2 amplification, indicating that SOX2 amplification may be an activating pathway to adenocarcinoma (40).
Another recent study showed that SOX2 is strongly and diffusely expressed in approximately 90% of pulmonary SCC and 20% of adenocarcinoma (41). When SOX2 expression was examined in stage I lung adenocarcinoma patients was detected in 50% of cases and it was more frequent in tumors from older and male patients (41). Compared to SOX2-negative tumors, SOX2 expression predicted a shorter time to tumor progression and shorter overall survival and appeared to be an independent predictor of poor outcome in stage I lung adenocarcinomas which may help stratify patients at increased risk for recurrence (41). Taken together, these results might suggest a prognostic role for SOX2 in lung adenocarcinomas. However, given the overall low frequency of SOX2 amplification and overexpression, the significance of this finding needs further evaluation.
As previously described, SOX2 gene amplification is more common in the SCCs of smokers while the incidence of SOX2 amplification is in the early stage of tumorigenesis in NSCLC. However SOX2 is also activated in more advanced SCC tumors (26,29). Therefore the SOX2 gene is not only activated by amplification but is also affected by other regulators that promote its transcription, affecting its downstream genes. SP cells isolated from established human NSCLC cell lines and tumors are highly enriched in NSCLC-CSCs and EGFR-Src-AKT signaling axis contributes significantly to the self-renewal of SP cells (11). Interestingly, SOX2 transcription factor is the predominant downstream target of EGFR signaling in these cells and plays a major role in self-renewal growth and expansion of SP cells, independent of Oct4 and Nanog (11).
EGFR tyrosine kinase inhibitors are able to downregulate self-renewal and SP phenotype. Singh et al., have reported that blocking EGF-receptors results in a significant decrease in SP frequency in both A549 and H1650 cells along with decreased EGFR phosphorylation as well as ABCG2 expression in both cell lines (11). Depletion of EGFR expression by erlotinib or gefitinib inhibits the self-renewal of SP cells and so does the combination of gefitinib with the irreversible EGFR-tyrosine kinase inhibitor BIBW2992 in the H1975 cell line with acquired resistance to gefitinib or erlotinib due to the secondary point mutation in exon 20 of EGFR (T790M) (11). When the downstream signaling events of EGFR expression were examined, it was found that c-Src, ERK and AKT signaling impinge transcription factors associated with stemness (Figure 1A) (11). EGFR inhibition by gefitinib or BIBW as well as inhibition of Src activity by dasatinib markedly decreased phosphorylation of EGFR, Src, ERK and AKT and reduced SOX2 expression; Oct4 and Nanog levels were not affected (Figure 1B,C) (11). The contribution of ERK and AKT pathways to EGFR mediated induction of SOX2 has also been examined. Phosphorylation of ERK is suppressed by the MEK inhibitor PD98059 and AKT phosphorylation is suppressed by the PI3-kinase inhibitor, LY294002. PI3-kinase inhibitors can also slightly inhibit ERK phosphorylation (Figure 1D) (11). However, as shown in Figure 1E, inhibition of MEK activity does not affect the levels of SOX2 while the PI3-kinase inhibition, markedly reduces its levels with corresponding reduction in SP frequency (11). Therefore, relative SOX2 expression and functions within the tumor-CSCs may be a major determinant in EGFR-targeted therapy against NSCLCs. This information might also be potentially useful to overcome the acquired resistance to EGFR therapies, by targeting downstream targets of EGFR signaling, including SOX2.
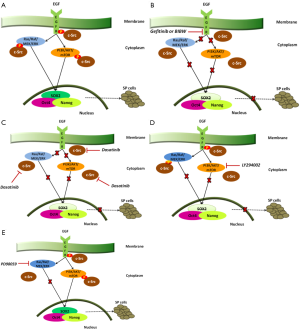
In summary, SOX2 might be a molecular target of lung adenocarcinomas. Transient transfection of SOX2 siRNA completely abrogated the tumorigenicity of SP cells in a lung adenocarcinoma cell line (LHK2) (42). SOX2 has a role in maintenance of stemness and tumorigenicity of human lung adenocarcinoma CSCs but further molecular analysis especially upstream and downstream of SOX2 should reveal the mechanisms of its tumorigenicity, making SOX2 a potential target for treatment.
Conclusions
SOX2 has been shown as hall mark of lung cancer but its role in lung cancer formation or progression has been partially elucidated. Amplification and overexpression of SOX2 are strongly associated with SCC morphology and favorable clinicopathological features in SCCs, including longer overall survival. In contrast, both events are less frequent in SCLC and rare in adenocarcinoma and of uncertain prognostic significance. The finding of SOX2 amplification/upregulation being frequent in lung SCCs, but rare in lung adenocarcinomas might reflect a fundamental molecular difference in carcinogenesis between these tumor entities.
The elucidation of SOX2-dependent pathways may identify novel therapeutic vulnerabilities in lung cancer and may uncover additional common pathways between cancer, normal development and the maintenance of pluripotency. The appearance of compensatory mechanisms favoring survival of cancer cells after therapy represents a limitation in therapies targeting EGFR and understanding but also overcoming EGFR-TKI resistance mechanisms in NSCLC patients has become a burning issue lately. Molecular pathways are interconnected, and thus, combination therapy is emerging as an appropriate strategy to treat those patients. Unfortunately very few patients undergo repeated tumor biopsies at the time when resistance develops to help guide appropriate therapeutic choices and the need to develop noninvasive methods to identify resistance mechanisms becomes more evident. For instance we should consider the possibility of using quantitative reverse transcriptase-PCR in measuring plasma SOX2 mRNA in lung cancer patients with gained-resistance to EGFR tyrosine kinase inhibitors and confirm whether high circulating plasma mRNA levels of SOX2 could be undocumented as a mechanism of resistance to EGFR-targeted therapy. In parallel the plasma mRNA measurement of the druggable downstream targets of EGFR signaling that regulate SOX2, can be also of great significance. Moreover SOX2 amplification may be more effectively identified by examining copy number changes by FISH specifically on individual circulating tumor cells. The recent advances in isolating circulating tumor cells suggest that this may be possible and can be combined with genotyping studies to examine mechanisms of resistance to EGFR tyrosine kinase inhibitors.
Except from targeting downstream targets of EGFR signaling that regulate SOX2 as mentioned above, being able to target SOX2 itself and other transcription factors involved in tumor initiation and maintenance can provide a unique opportunity for anti-cancer intervention. However, because of their lack of small molecule binding pockets, transcription factors are currently an example of ‘undruggable targets’. Thus, novel strategies to effectively down-regulate these targets are required. Recently Zinc-finger-based artificial transcription factors (ATFs) were able to reactivate the expression of the tumor-suppressor genes and repress potential oncogenes including SOX2 in breast cancer cell lines (43). These data suggest that the targeted down-regulation of highly expressed oncogenes using ATF-based technologies can be used as a powerful tool for the long-term targeting of oncogenic TFs with potential application in cancer biology. In summary, the above data elucidate and offer novel perspectives on the multiple roles that the transcription factor SOX2 exerts on carcinogenesis. SOX2 that is expressed in lung SCC and adenocarcinoma, but also in SCLC tissues can act as novel unite marker and ideal therapeutic target. In view of the fact that the transcriptional activity of SOX2 is critical in mediating tumorigenesis, we believe that further studies investigating how SOX2 activity is regulated will be highly worthwhile.
Acknowledgements
Disclosure: The authors declare no conflict of interest.
References
- Kiefer JC. Back to basics: Sox genes. Dev Dyn 2007;236:2356-66. [PubMed]
- Bylund M, Andersson E, Novitch BG, et al. Vertebrate neurogenesis is counteracted by Sox1-3 activity. Nat Neurosci 2003;6:1162-8. [PubMed]
- Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006;126:663-76. [PubMed]
- Zhou S, Schuetz JD, Bunting KD, et al. The ABC transporter Bcrp1/ABCG2 is expressed in a wide variety of stem cells and is a molecular determinant of the side-population phenotype. Nat Med 2001;7:1028-34. [PubMed]
- Hirschmann-Jax C, Foster AE, Wulf GG, et al. A distinct “side population” of cells with high drug efflux capacity in human tumor cells. Proc Natl Acad Sci U S A 2004;101:14228-33. [PubMed]
- Kruger JA, Kaplan CD, Luo Y, et al. Characterization of stem cell-like cancer cells in immune-competent mice. Blood 2006;108:3906-12. [PubMed]
- Seo E, Basu-Roy U, Zavadil J, et al. Distinct functions of Sox2 control self-renewal and differentiation in the osteoblast lineage. Mol Cell Biol 2011;31:4593-608. [PubMed]
- Okumura-Nakanishi S, Saito M, Niwa H, et al. Oct-3/4 and Sox2 regulate Oct-3/4 gene in embryonic stem cells. J Biol Chem 2005;280:5307-17. [PubMed]
- Sinner D, Rankin S, Lee M, et al. Sox17 and beta-catenin cooperate to regulate the transcription of endodermal genes. Development 2004;131:3069-80. [PubMed]
- Chen Y, Shi L, Zhang L, et al. The molecular mechanism governing the oncogenic potential of SOX2 in breast cancer. J Biol Chem 2008;283:17969-78. [PubMed]
- Singh S, Trevino J, Bora-Singhal N, et al. EGFR/Src/Akt signaling modulates Sox2 expression and self-renewal of stem-like side-population cells in non-small cell lung cancer. Mol Cancer 2012;11:73. [PubMed]
- Li XL, Eishi Y, Bai YQ, et al. Expression of the SRY-related HMG box protein SOX2 in human gastric carcinoma. Int J Oncol 2004;24:257-63. [PubMed]
- Gangemi RM, Griffero F, Marubbi D, et al. SOX2 silencing in glioblastoma tumor-initiating cells causes stop of proliferation and loss of tumorigenicity. Stem Cells 2009;27:40-8. [PubMed]
- Lu Y, Futtner C, Rock JR, et al. Evidence that SOX2 overexpression is oncogenic in the lung. PLoS One 2010;5:e11022. [PubMed]
- Leung EL, Fiscus RR, Tung JW, et al. Non-small cell lung cancer cells expressing CD44 are enriched for stem cell-like properties. PLoS One 2010;5:e14062. [PubMed]
- Rudin CM, Durinck S, Stawiski EW, et al. Comprehensive genomic analysis identifies SOX2 as a frequently amplified gene in small-cell lung cancer. Nat Genet 2012;44:1111-6. [PubMed]
- Bae KM, Su Z, Frye C, et al. Expression of pluripotent stem cell reprogramming factors by prostate tumor initiating cells. J Urol 2010;183:2045-53. [PubMed]
- Xu XL, Xing BC, Han HB, et al. The properties of tumor-initiating cells from a hepatocellular carcinoma patient’s primary and recurrent tumor. Carcinogenesis 2010;31:167-74. [PubMed]
- Basu-Roy U, Seo E, Ramanathapuram L, et al. Sox2 maintains self renewal of tumor-initiating cells in osteosarcomas. Oncogene 2012;31:2270-82. [PubMed]
- Lengerke C, Fehm T, Kurth R, et al. Expression of the embryonic stem cell marker SOX2 in early-stage breast carcinoma. BMC Cancer 2011;11:42. [PubMed]
- van Meerbeeck JP, Fennell DA, De Ruysscher DK. Small-cell lung cancer. Lancet 2011;378:1741-55. [PubMed]
- Jemal A, Siegel R, Xu J, et al. Cancer statistics, 2010. CA Cancer J Clin 2010;60:277-300. [PubMed]
- Chen S, Xu Y, Chen Y, et al. SOX2 gene regulates the transcriptional network of oncogenes and affects tumorigenesis of human lung cancer cells. PLoS One 2012;7:e36326. [PubMed]
- Gontan C, de Munck A, Vermeij M, et al. Sox2 is important for two crucial processes in lung development: branching morphogenesis and epithelial cell differentiation. Dev Biol 2008;317:296-309. [PubMed]
- Güre AO, Stockert E, Scanlan MJ, et al. Serological identification of embryonic neural proteins as highly immunogenic tumor antigens in small cell lung cancer. Proc Natl Acad Sci U S A 2000;97:4198-203. [PubMed]
- Sholl LM, Long KB, Hornick JL. Sox2 expression in pulmonary non-small cell and neuroendocrine carcinomas. Appl Immunohistochem Mol Morphol 2010;18:55-61. [PubMed]
- Maier S, Wilbertz T, Braun M, et al. SOX2 amplification is a common event in squamous cell carcinomas of different organ sites. Hum Pathol 2011;42:1078-88. [PubMed]
- Bass AJ, Watanabe H, Mermel CH, et al. SOX2 is an amplified lineage-survival oncogene in lung and esophageal squamous cell carcinomas. Nat Genet 2009;41:1238-42. [PubMed]
- Hussenet T, Dali S, Exinger J, et al. SOX2 is an oncogene activated by recurrent 3q26.3 amplifications in human lung squamous cell carcinomas. PLoS One 2010;5:e8960. [PubMed]
- Yuan P, Kadara H, Behrens C, et al. Sex determining region Y-Box 2 (SOX2) is a potential cell-lineage gene highly expressed in the pathogenesis of squamous cell carcinomas of the lung. PLoS One 2010;5:e9112. [PubMed]
- Sasaki H, Yokota K, Hikosaka Y, et al. Increased Sox2 copy number in lung squamous cell carcinomas. Exp Ther Med 2012;3:44-48. [PubMed]
- Hussenet T, du Manoir S. SOX2 in squamous cell carcinoma: amplifying a pleiotropic oncogene along carcinogenesis. Cell Cycle 2010;9:1480-6. [PubMed]
- Toschi L, Finocchiaro G, Teresa T. SOX2 and FGFR1 gene copy number in surgically resected non-small cell lung cancer (NSCLC). J Clin Oncol 2012;30:abstr 7061.
- Ji H, Ramsey MR, Hayes DN, et al. LKB1 modulates lung cancer differentiation and metastasis. Nature 2007;448:807-10. [PubMed]
- Que J, Okubo T, Goldenring JR, et al. Multiple dose-dependent roles for Sox2 in the patterning and differentiation of anterior foregut endoderm. Development 2007;134:2521-31. [PubMed]
- McCaughan F, Pole JC, Bankier AT, et al. Progressive 3q amplification consistently targets SOX2 in preinvasive squamous lung cancer. Am J Respir Crit Care Med 2010;182:83-91. [PubMed]
- Wilbertz T, Wagner P, Petersen K, et al. SOX2 gene amplification and protein overexpression are associated with better outcome in squamous cell lung cancer. Mod Pathol 2011;24:944-53. [PubMed]
- Long KB, Hornick JL. SOX2 is highly expressed in squamous cell carcinomas of the gastrointestinal tract. Hum Pathol 2009;40:1768-73. [PubMed]
- Freier K, Knoepfle K, Flechtenmacher C, et al. Recurrent copy number gain of transcription factor SOX2 and corresponding high protein expression in oral squamous cell carcinoma. Genes Chromosomes Cancer 2010;49:9-16. [PubMed]
- Cai YR, Zhang HQ, Zhang ZD, et al. Detection of MET and SOX2 amplification by quantitative real-time PCR in non-small cell lung carcinoma. Oncol Lett 2011;2:257-264. [PubMed]
- Sholl LM, Barletta JA, Yeap BY, et al. Sox2 protein expression is an independent poor prognostic indicator in stage I lung adenocarcinoma. Am J Surg Pathol 2010;34:1193-8. [PubMed]
- Nakatsugawa M, Takahashi A, Hirohashi Y, et al. SOX2 is overexpressed in stem-like cells of human lung adenocarcinoma and augments the tumorigenicity. Lab Invest 2011;91:1796-804. [PubMed]
- Stolzenburg S, Rots MG, Beltran AS, et al. Targeted silencing of the oncogenic transcription factor SOX2 in breast cancer. Nucleic Acids Res 2012;40:6725-40. [PubMed]


