
PD-L1 detection on circulating tumor-derived extracellular vesicles (T-EVs) from patients with lung cancer
Introduction
Immunotherapies with immune checkpoint inhibitors (ICIs) have produced encouraging results in patients with non-small cell lung cancer (NSCLC) (1-6). Several antibodies against programmed cell death-1 (PD-1) and programmed cell death ligand-1 (PD-L1) have been approved as treatments in monotherapy or in combination with chemotherapy, other immunotherapies, and anti-angiogenesis agents in many countries (7-10). The magnitude of the clinical benefit is generally associated with the tumor PD-L1 expression level (1-5,11,12). One of the current approaches for the use of these therapeutic products is to stratify or select eligible patient populations by performing immunohistochemistry (IHC) with an anti-PD-L1 antibody on tumor tissues specimens. The expression of the PD-L1 protein is determined by calculating the tumor proportion score (TPS), which is the percentage of viable tumor cells showing partial or complete membrane staining at any intensity. The specimen is considered to express PD-L1 if the TPS ≥1% and to express PD-L1 at high levels if the TPS ≥50% (1,13). Although this approach remains controversial (14,15), patients with a higher PD-L1 TPS may obtain more benefits from ICIs therapies (1,7,13,16-18).
Although PD-L1 IHC is widely used by clinicians to identify patients eligible for immunotherapy, it has several limitations. The limited availability of tumor tissues for testing, the variation driven by the biopsy site, and the variation in the reading of the assays by pathologists hamper the use of IHC as an ideal system for assessing PD-L1 expression (19,20). In addition, the regulation of PD-L1 expression is a dynamic process that is unable to be adequately monitored by examining a tumor tissue biopsy. Thus, a less invasive, easier and more accurate technique to monitor tumor PD-L1 expression, such as the liquid biopsy, is urgently needed.
Tumor-derived extracellular vesicles (T-EVs), such as exosomes (30–150 nm) and microvesicles (150–1,000 nm), whose molecular and genetic contents partially resemble the tumor cells from which they originate, are currently viewed as promising “liquid biopsies”. T-EVs carry bioactive molecules and play diverse roles in tumor progression, including invasion, immune modulation, neovascularization, and metastasis (21-23). PD-L1 was reported to be expressed on the surface of EVs derived from tumor cell lines and detected in blood samples from patients with cancer (24-26). According to recent studies, PD-L1 on EVs contributes to suppressing anti-tumor immunity and has the potential to serve as a biomarker in patients with cancer (24,25,27). Additionally, some studies showed significant correlations between the levels of PD-L1 in EVs and the pathological features of patients with cancer, such as the tumor size, stage or treatment response (24-26,28). However, the existing assays and technologies to detect PD-L1+ EVs are either limited in sensitivity or highly complex, which prevent their use in medical practice. Thus, a method to rapidly detect and distinguish PD-L1+ EVs, particularly those derived from tumor cells, has yet to be developed.
Recently, using a single molecule array (Simoa), a highly sensitive immunoassay technology (29-32), we developed automatic EV/T-EV detection assays, enabling the direct profiling of EVs/T-EVs from the plasma of patients with cancer (33). In the current study, by capturing T-EVs with the same marker, epithelial cell adhesion molecule (Epcam), an important surface marker of epithelial tumor cell used as a biomarker of circulating tumor cells (CTCs) and T-EVs (34), we aim to develop a Simoa immunoassay prototype to detect PD-L1+ T-EVs from blood samples of patients with lung cancer and to compare PD-L1 expression in T-EVs and tumor tissues.
We present the following article in accordance with the MDAR reporting checklist (available at http://dx.doi.org/10.21037/tlcr-20-1277).
Methods
Ethical statement
The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by institutional review board of Fudan University Shanghai Cancer Center (No.: 050432-4-1911D) and individual consent for this retrospective analysis was waived.
Prototype of the Simoa assay for detecting EVs
Antibodies against Epcam and PD-L1 were selected and prepared for capture and detection according to the manufacturer’s protocol (Quanterix, Lexington, MA, USA). Bead conjugation protocol was followed by an incubation at 4 °C. The capture antibody concentration was adjusted to 0.2 mg/mL with Bead Conjugation Buffer and then paramagnetic carboxylated microparticles (Quanterix) were activated with 0.3 mg/mL 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride (EDC) (Thermo Fisher Scientific, Waltham, MA, USA). Three microliters of the biotin solution (2 mg of NHS-PEG4-Biotin dissolved in 383 µL of ddH2O) were added to 100 µL of the detection antibody solution (1.0 mg/mL) for a molar biotinylation ratio of 40× to start the biotinylation reaction. The concentration of the recovered antibody was adjusted to 0.2 mg/mL and stored at 4 °C.
In order to screen the best antibody pair for Simoa prototype, three monoclonal antibodies of PD-L1 were purchased from Abcam and Origene (Table S1). An antibody for Epcam (MAB9601, R&D Systems) confirmed in a previously study (33) was taken as the capture antibody. Exosomes were collected from cell culture supernatant of HCT-116 for antibody pair testing due to its positive expression of Epcam and PD-L1 (35,36). Finally, the antibody pair of MAB9601-TA507087 gave the highest signal/background ratio among all the antibodies tested, thus was selected for the further study (Figure S1).
Simoa assay setup
All Simoa measurements were performed using a fully automated Simoa HD-1 Analyzer (Quanterix). The microparticles coated with the Epcam capture antibody were diluted in Bead Diluent (Quanterix) to 500,000 microparticles per test. The PD-L1 detection antibody was diluted in Homebrew Detector/Sample Diluent (Quanterix) to a working concentration of 2 µg/mL. The streptavidin-β-galactosidase concentrate was diluted to a working concentration of 150 pM. The assay includes 3 steps. First, 25 µL of the microparticle solution and 100 µL of patient plasma (1:4 diluted) were incubated for 45 minutes in a reaction cuvette (Quanterix). Second, after several washes, the mixture and PD-L1 detection antibody were incubated for 5 minutes 15 seconds. Finally, 100 µL of Streptavidin-β-galactosidase (SBG) were added and incubated for 5 minutes 15 seconds. Resorufin β-D-galactopyranoside (RGP) was added after several washes and the sample was loaded into the array. The array was then sealed with oil and the microparticles were imaged. The automated analysis was performed using HD-1 Analyzer software (Quanterix). Simoa signal is expressed in average per bead (AEB) as previous described (31,33). In short, AEB is determined by counting the number of wells containing both a bead and fluorescent signal (“on” well) relative to the total number of wells containing beads, using Poisson statistics and the digital or analog methods based on high or low concentrations of captured analyte. At low concentrations, the ratio of analytes to beads is small resulting in statistical distribution of individual molecules on the beads, giving Simoa its single molecule sensitivity.
Cell culture and plasma
Two lung cancer cell lines (A549, ATCC® CCL-185™ and SK-MES1, ATCC® HTB-58™) were cultured according to a standard protocol at Fudan University Shanghai Cancer Center-Institut Mérieux Laboratory. Cells were cultured in DMEM (L120KJ, YuanPei Biotech) supplemented with 10% FBS (SH30084.03, HyClone) until the cells reached 80% culture confluence, and then washed twice with DMEM. After 36 hours of culture in DMEM, culture media were collected for direct testing or EV purification. For IFNγ stimulation, cells were treated with 10 ng/mL IFNγ (R&D Systems) for 36 hours before collecting the culture media.
Clinical samples analyzed by Simoa platform were EDTA plasma samples collected from lung cancer patients before surgery. Three milliliter of patient whole blood were collected into the EDTA tube. Plasma was separated within 2 hours under 3,000 rpm centrifugation and transferred to the tissue bank of Fudan University Shanghai Cancer Center (FUSCC). The plasma samples used in this study were mainly collected in 2018 and stored at −80 °C. Each participant signed an informed consent form allowing their samples to be used for research activities before they were enrolled in this study.
Exosome/microvesicle purification
Differential ultra-centrifugation was used to purify exosomes as described previously (37). Briefly, after collecting cell culture media, a low speed centrifugation (300 g, 10 minutes) step was applied to remove dead cells. Then, another centrifugation step at 2,000 g for 20 minutes was performed to remove cell debris. A subsequent 40-minute centrifugation step at 10,000 g was applied to remove microvesicles. Exosomes were then collected by ultracentrifugation at 100,000 g for 2 hours using a Beckman Coulter Optima™ XP ultracentrifuge. Pellets were washed once with PBS and ulracentrifuged again at 100,000 g for 2 hours. The exosome concentration was calculated using a Flow NanoAnalyzer (NanoFCM Inc., Xiamen, China) according to the manufacturer’s instructions.
Exosome flow cytometry
An on-bead flow cytometry is used to detect exosomes/EVs (38,39). In brief, one hundred microliters of the exosome solution were mixed with 10 µL of 4 µm aldehyde/sulfate latex beads (S37225, Thermo Fisher) for 15 minutes at room temperature with continuous rotation. After adding 1 mL of PBS, the mixture was incubated for 30 minutes at room temperature in rotation mode and centrifuged; then, the supernatant was removed. The reaction was stopped by incubating it with Stop Buffer (1 mM glycine, 2% BSA, 1× PBS) for 30 minutes. After 3 washes, exosomes bound to beads were blocked with 1 mL of 10% BSA for 30 minutes. Following 3 washes, the beads were sequentially incubated with 1 µg of the PD-L1 antibody (clone 28-8, ab205921, Abcam) and the HRP-Goat Anti-Rabbit IgG H&L (ab97051) secondary antibody (1:20,000 dilution).
Transmission electronic microscopy
Electronic microscopy was performed at the Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences (Shanghai). Freshly isolated exosomes were layered on copper grids with 0.125% Formvar in chloroform and stained with 1% uranyl acetate in ddH2O. Immediately, exosomes on the grids were visualized using a JEOL JEM-1011 transmission electron microscope (TESCAN).
Analysis of PD-L1 protein expression using IHC
IHC staining for the PD-L1 protein was performed on formalin-fixed paraffin-embedded (FFPE) tumor sections (4 µm thick) immediately after the operation, as described previously (40). In specimens with a maximum diameter greater than 1 cm, a representative slide was selected for staining. The representative slide was defined as a section that contained the most diverse histological subtypes. This assay was performed on the Dako Autostainer Link 48 platform with an automated staining protocol using a mouse monoclonal anti-PD-L1 antibody (22C3). PD-L1 expression was evaluated by 2 board-certified pathologists in FUSCC, who were blinded to clinical data and patient outcomes, via calculating the TPS, which is defined as the percentage of PD-L1-positive tumor cells (TCs) relative to the total number of TCs. The evaluation of the score included partial or complete membranous staining (at least a 1+ intensity). All other cells, such as tumor-associated immune cells, normal/non-neoplastic cells, and necrotic cells, were excluded from the evaluation. PD-L1 expression on TCs was classified into three levels: negative expression (TPS <1%), low expression (TPS 1–49%), and high expression (TPS ≥50%).
Statistical analysis
Plasma levels of circulating Epcam-PD-L1 EVs were compared with PD-L1 expression in solid tumor (TPS). A ROC curve was generated to calculate the area under curve (AUC), sensitivity and specificity of the Simoa Epcam-PD-L1 assay in diagnosing PD-L1 positive patients. The highest AUC was determined by screening all possible TPS cutoffs. The tumor volume was calculated as the product of the three dimensions of the tumor in pathological records. Spearman’s correlation coefficient between the Simoa Epcam-PD-L1 level, TPS and TPS multiplied by tumor volume were calculated to study their associations.
Results
Design of the EV detection model
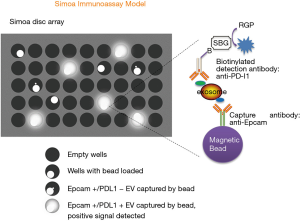
The objective of this study was to detect PD-L1 expression levels on circulating tumor-derived EVs. Therefore, a Simoa EV detection assay was designed, as shown in Figure 1. In the Simoa disc array, over 20,000 wells were used in the signal collection step, with diameters of 4.5 µm and depths of 3.25 µm. Considering the diameter of magnetic beads (2.7 µm diam.), the Simoa technology is capable of detecting most EVs, including exosomes (30–150 nm). First, a sandwich antibody complex is formed on microscopic beads. In samples containing EVs with Epcam expression, EVs are captured by Epcam capture antibodies on the magnetic beads. If EVs also express PD-L1, fluorophores are generated and fluorescence images are captured to calculate the signals. Several antibodies against Epcam and PD-L1 were tested based on the development of the homemade Simoa kit; Epcam (MAB9601, R&D Systems) and PD-L1 (TA507087, Origene) antibodies were selected for our Simoa prototype.
Validation of the Epcam-PD-L1 Simoa assay
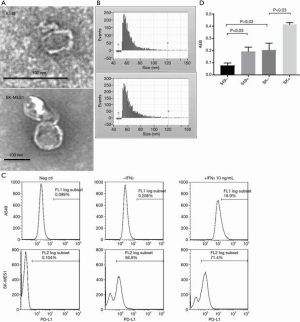
Exosomes were isolated from two lung cancer cell lines, A549 and SK-MES1, using a previously described ultracentrifugation method to further validate the PD-L1+ EV detection prototype (37). Isolated exosomes were further analyzed using transmission electron microscopy (TEM) and showed the expected round morphology (Figure 2A). The concentration and size distribution of exosomes were analyzed using a Flow NanoAnalyzer (Figure 2B) (41). Based on the EV flow cytometry results, PD-L1 was almost undetectable on the surface of exosomes isolated from A549 cell line, while high PD-L1 expression was observed on the exosomes from the SK-MES1 cell line (Figure 2C, left panel). IFNγ treatment has been reported to stimulate the upregulation of PD-L1 on the surface of tumor cells and EVs from various cancer cells (21,22). After IFNγ activation, PD-L1 expression levels increased in exosomes from both cell lines (Figure 2C, right panel). Meanwhile, the same samples were analyzed using the Simoa PD-L1-EV assay. Consistent with the flow cytometry results, Simoa testing showed higher signals in SK-MES1 cells than in A549 cells, and the IFNγ treatment increased PD-L1 expression in both cell lines (Figure 2D). Thus, the Simoa Epcam-PD-L1 prototype may be a good assay to measure PD-L1+ exosomes.
Association between Simoa Epcam-PD-L1 level and the PD-L1 TPS
We next tested Simoa Epcam-PD-L1 assay on plasma samples from 35 patients with lung cancer. Clinicopathological characteristics of the patients are summarized in Table 1. The median age of the patients was 64 years (range, 45–76 years). Twenty-four of 35 (68.6%) patients were male. More than half (65.7%) of the patients had tumor smaller than 3 cm in largest dimension. Approximately 45.7% (16/35) of the patients presented with lymph node metastasis. TNM stage showed that more T1 (42.9%) and non-metastasis (91.4%) patients were enrolled. Vascular invasion was observed in 65.7% (23/35) of patients. Clinical information for the TPS of PD-L1 IHC staining (22C3 PharmDx, Agilent Technologies, Santa Clara, CA, USA) was available for all patients. Twenty-eight patients had a TPS ≥1%, which is the clinical accepted cutoff for a PD-L1-expressing patient (13).

Full table
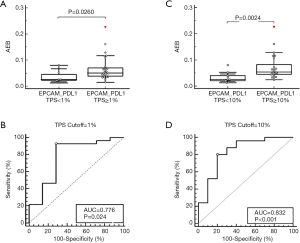
We therefore compared the Simoa Epcam-PD-L1 levels between PD-L1-expressing and non-expressing patients using the aforementioned TPS cutoff (Figure 3A). When the TPS cutoff was set to 1%, the Simoa Epcam-PD-L1 values were significantly higher in positive samples (P=0.026, Figure 3A), with an AUC of 0.776, a sensitivity of 92.86% and a specificity of 71.43% at the highest Youden index (Figure 3B). The highest AUC was observed when the TPS cutoff was set to 10%, with 25 positive (TPS ≥10%) and 10 negative (TPS <10%) samples. At this cutoff, the Simoa Epcam-PD-L1 level was significantly increased in positive samples (P=0.0024, Figure 3C). The AUC reached 0.832, with both a sensitivity and specificity of 80% at the highest Youden index (Figure 3D). It has been demonstrated that TPS ≥50% is for PD-L1 high expression patients who would be response positive to ICIs treatment. We therefore also evaluated a discrimination of the Epcam-PD-L1 expression on these PD-L1 high expression patients. At 50% cutoff, Epcam-PD-L1 showed an increase in positive samples but not statistically significant (P=0.109), AUC was at 0.661 with sensitivity at 100% specificity at 36.84% at the highest Youden index (Figure S2).
In addition, we compared the Simoa results to the TPS score for each patient. Spearman’s correlation coefficient for Simoa Epcam-PD-L1 and TPS was equal to 0.428 (P=0.0104, 95% CI 0.110 to 0.666, Figure 4A). A small increase in Spearman’s correlation coefficient (0.482, P=0.003) was observed when the TPS was multiplied by the tumor volume (Figure 4B).
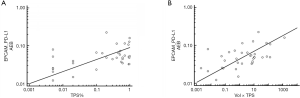
Based on our results, PD-L1+ EV/exosome levels measured using the Simoa Epcam-PD-L1 assay are significantly correlated with the TPS.
Discussion
Although still controversial, PD-L1 IHC is still one of the putative predictors of the response to PD-1/PD-L1-targeted checkpoint inhibitors. Since 2015, high expression of the PD-L1 protein in tumor cells or tumor microenvironment (TME) has been identified to be a logical biomarker for predicting the efficacy of ICI therapy and was approved by the U.S. Food and Drug Administration as an indicator of initiating treatment for various solid tumors (15). On the other hand, the detection of PD-L1 expression through liquid biopsy, including tumor-derived soluble PD-L1 (sPD-L1) and PD-L1 on CTC or EVs/exosomes has garnered increasing attention to date. However, no evidence is available on whether circulating PD-L1 expression was consistent with tissue PD-L1 expression, and whether circulating PD-L1 levels had a similar value to predict the tumor response as tissue PD-L1 expression. In addition, the identification and quantification of biomarkers on EVs/exosomes in clinical samples remains challenging due to the complex isolation process. For example, to evaluate exosomes by the flow cytometry technology requires isolation of exosomes before detection, which is not feasible in clinical setting. Due to the small size of EVs/exosomes, an aldehyde/sulfate latex beads need to be used in the conventional flow cytometry to capture exosomes but in a non-specific way (37,38). In contrast, Simoa platform might provide an ultrasensitive, non-invasive, fully automated, and high-throughput EV detection assay with double EVs biomarkers targeting (33). In the current study, for the first time, we developed a PD-L1+ EV detection assay based on the Simoa technology and identified a significant correlation of PD-L1 expression between T-EVs and tumor tissues. According to our results, our Simoa prototype might provide a non-invasive and dynamic method to monitor tumor PD-L1 expression in patients undergoing ICI therapy.
In our study, plasma levels of PD-L1+ T-EVs were significantly increased in PD-L1-positive patients (TPS ≥1%) compared to PD-L1-negative patients (TPS <1%) (P=0.026). When the 10% threshold for the TPS was used to distinguish PD-L1 positive patients, the best performance of the Epcam-PD-L1 assay was observed (AUC =0.832). At TPS cutoff set as 50%, the performance of Epcam-PD-L1 assay decreased to AUC =0.661. Based on those results, more PD-L1 IHC-positive patients also present high PD-L1 signals on T-EVs, while relatively low levels of PD-L1+ T-EVs are detected in PD-L1 IHC-negative patients. This characteristic might be useful in the future for monitoring therapeutic responses to ICIs. Moreover, the Simoa automatic exosome detection system potentially represents a nice technology to use in clinic practice.
Although the Simoa assay and TPS showed a similar trend in PD-L1 expression between PD-L1-positive and -negative patient groups, a subsequent analysis of Spearman’s correlation coefficients between the results obtained with the two methods showed that the association between the PD-L1+ T-EV signal and PD-L1-IHC TPS results was still weak (R=0.428, P=0.0104). We postulated that the discrepancy between the two methods might be explained by several factors. First, inter-tumor heterogeneity in PD-L1 expression is an important issue for an accurate TPS, while circulating T-EVs might theoretically provide a better representation of the whole tumor. The TPS defines the portion of tumor cells in the observed slide. Thus, the multiplication of the TPS by the tumor volume may be a good method to obtain an approximate number of tumor cells. Because we detected circulating exosomes in plasma with our Simoa Epcam-PD-L1 assay, this absolute tumor cell burden may be more relevant. Indeed, when TPS values are multiplied by the tumor volume, the correlation with the PD-L1+ T-EVs increased (Figure 4B, R=0.482, P=0.003).
Second, although Epcam may be the best biomarker for T-EVs and was chosen to capture T-EVs in our Simoa prototype, it is not expressed in 100% of carcinomas (33). High-level and mostly homogenous expression of Epcam were observed on 85% of adenocarcinomas and on 72% of squamous cell carcinomas (34). Finally, the well-known PD-L1 IHC antibodies, such as 22C3 (Dako), 28-8 (Dako), SP142 (Ventana) and SP263 (Ventana), show different efficiencies for PD-L1 tissue staining and therefore different cutoffs for PD-L1-positive expression must be used (20). In our study, the PD-L1 TPS results were obtained with the 22C3 antibody, while an Origene antibody (TA507086) was chosen for the Simoa prototype due to its good performance. The differences in the efficiencies of the antibodies used in the two methods might also be responsible for the variation observed when comparing the results.
The encouraging results obtained with the Simoa PD-L1+T-EVs assay were based on a population with a limited size. The current results must now be confirmed in a larger patient cohort. Additionally, other assays might be performed to obtain a better understanding of the technical issues raised above, including Epcam specificity and the different effects of anti-PD-L1 antibodies and finally to standardize procedures before clinical usage. At last, additional clinical trials should be conducted to determine whether PD-L1 expression on the circulating T-EVs has a similar value to tissue PD-L1 IHC in predicting the tumor response to ICI therapies and its expression cutoff sensitive to ICIs therapy should also be evaluated.
Acknowledgments
Funding: This work was supported by the National Science Foundation of China (grant numbers 81972185), Shanghai Natural Science Foundation (grant numbers 18ZR1407800), Rising-Star Program (grant number19QA1402200).
Footnote
Reporting Checklist: The authors have completed the MDAR reporting checklist. Available at http://dx.doi.org/10.21037/tlcr-20-1277
Data Sharing Statement: Available at http://dx.doi.org/10.21037/tlcr-20-1277
Peer Review File: Available at http://dx.doi.org/10.21037/tlcr-20-1277
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tlcr-20-1277). PW reports that she has received research grants from National Science Foundation of China, Shanghai Natural Science Foundation of China, and Rising-Star Program of China. The other authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by institutional review board of Fudan University Shanghai Cancer Center (No.: 050432-4-1911D) and individual consent for this retrospective analysis was waived.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Herbst RS, Baas P, Kim DW, et al. Pembrolizumab versus docetaxel for previously treated, PD-L1-positive, advanced non-small-cell lung cancer (KEYNOTE-010): a randomised controlled trial. Lancet 2016;387:1540-50. [Crossref] [PubMed]
- Petrylak DP, Powles T, Bellmunt J, et al. Atezolizumab (MPDL3280A) Monotherapy for Patients With Metastatic Urothelial Cancer: Long-term Outcomes From a Phase 1 Study. JAMA Oncol 2018;4:537-44. [Crossref] [PubMed]
- Fehrenbacher L, Spira A, Ballinger M, et al. Atezolizumab versus docetaxel for patients with previously treated non-small-cell lung cancer (POPLAR): a multicentre, open-label, phase 2 randomised controlled trial. Lancet 2016;387:1837-46. [Crossref] [PubMed]
- Borghaei H, Paz-Ares L, Horn L, et al. Nivolumab versus Docetaxel in Advanced Nonsquamous Non-Small-Cell Lung Cancer. N Engl J Med 2015;373:1627-39. [Crossref] [PubMed]
- Rizvi NA, Mazieres J, Planchard D, et al. Activity and safety of nivolumab, an anti-PD-1 immune checkpoint inhibitor, for patients with advanced, refractory squamous non-small-cell lung cancer (CheckMate 063): a phase 2, single-arm trial. Lancet Oncol 2015;16:257-65. [Crossref] [PubMed]
- Copur MD, Gauchan D, Ramaekers R. Durvalumab in Stage III Non-Small-Cell Lung Cancer. N Engl J Med 2018;378:868. [Crossref] [PubMed]
- Gandhi L, Rodriguez-Abreu D, Gadgeel S, et al. Pembrolizumab plus Chemotherapy in Metastatic Non-Small-Cell Lung Cancer. N Engl J Med 2018;378:2078-92. [Crossref] [PubMed]
- Brahmer JR, Lacchetti C, Schneider BJ, et al. Management of Immune-Related Adverse Events in Patients Treated With Immune Checkpoint Inhibitor Therapy: American Society of Clinical Oncology Clinical Practice Guideline. J Clin Oncol 2018;36:1714-68. [Crossref] [PubMed]
- Haanen JB, Carbonnel F, Robert C, et al. Management of toxicities from immunotherapy: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol 2017;28:iv119-42. [Crossref] [PubMed]
- Haanen JB, Carbonnel F, Robert C, et al. Management of toxicities from immunotherapy: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol 2018;29:iv264-6. [Crossref] [PubMed]
- Herbst RS, Soria JC, Kowanetz M, et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature 2014;515:563-7. [Crossref] [PubMed]
- Ibrahim R, Stewart R, Shalabi A. PD-L1 blockade for cancer treatment: MEDI4736. Semin Oncol 2015;42:474-83. [Crossref] [PubMed]
- Mok TSK, Wu YL, Kudaba I, et al. Pembrolizumab versus chemotherapy for previously untreated, PD-L1-expressing, locally advanced or metastatic non-small-cell lung cancer (KEYNOTE-042): a randomised, open-label, controlled, phase 3 trial. Lancet 2019;393:1819-30. [Crossref] [PubMed]
- Patel SP, Kurzrock R. PD-L1 Expression as a Predictive Biomarker in Cancer Immunotherapy. Mol Cancer Ther 2015;14:847-56. [Crossref] [PubMed]
- Sul J, Blumenthal GM, Jiang X, et al. FDA Approval Summary: Pembrolizumab for the Treatment of Patients With Metastatic Non-Small Cell Lung Cancer Whose Tumors Express Programmed Death-Ligand 1. Oncologist 2016;21:643-50. [Crossref] [PubMed]
- Reck M, Rodriguez-Abreu D, Robinson AG, et al. Updated Analysis of KEYNOTE-024: Pembrolizumab Versus Platinum-Based Chemotherapy for Advanced Non-Small-Cell Lung Cancer With PD-L1 Tumor Proportion Score of 50% or Greater. J Clin Oncol 2019;37:537-46. [Crossref] [PubMed]
- Paz-Ares L, Luft A, Vicente D, et al. Pembrolizumab plus Chemotherapy for Squamous Non-Small-Cell Lung Cancer. N Engl J Med 2018;379:2040-51. [Crossref] [PubMed]
- West H, McCleod M, Hussein M, et al. Atezolizumab in combination with carboplatin plus nab-paclitaxel chemotherapy compared with chemotherapy alone as first-line treatment for metastatic non-squamous non-small-cell lung cancer (IMpower130): a multicentre, randomised, open-label, phase 3 trial. Lancet Oncol 2019;20:924-37. [Crossref] [PubMed]
- Remon J, Chaput N, Planchard D. Predictive biomarkers for programmed death-1/programmed death ligand immune checkpoint inhibitors in nonsmall cell lung cancer. Curr Opin Oncol 2016;28:122-9. [Crossref] [PubMed]
- Hirsch FR, McElhinny A, Stanforth D, et al. PD-L1 Immunohistochemistry Assays for Lung Cancer: Results from Phase 1 of the Blueprint PD-L1 IHC Assay Comparison Project. J Thorac Oncol 2017;12:208-22. [Crossref] [PubMed]
- Kogure T, Yan IK, Lin WL, et al. Extracellular Vesicle-Mediated Transfer of a Novel Long Noncoding RNA TUC339: A Mechanism of Intercellular Signaling in Human Hepatocellular Cancer. Genes Cancer 2013;4:261-72. [Crossref] [PubMed]
- Atay S, Godwin AK. Tumor-derived exosomes: A message delivery system for tumor progression. Commun Integr Biol 2014;7:e28231 [Crossref] [PubMed]
- Weston WW, Ganey T, Temple HT. The Relationship between Exosomes and Cancer: Implications for Diagnostics and Therapeutics. BioDrugs 2019;33:137-58. [Crossref] [PubMed]
- Ricklefs FL, Alayo Q, Krenzlin H, et al. Immune evasion mediated by PD-L1 on glioblastoma-derived extracellular vesicles. Sci Adv 2018;4:eaar2766 [Crossref] [PubMed]
- Chen G, Huang AC, Zhang W, et al. Exosomal PD-L1 contributes to immunosuppression and is associated with anti-PD-1 response. Nature 2018;560:382-6. [Crossref] [PubMed]
- Theodoraki MN, Yerneni SS, Hoffmann TK, et al. Clinical Significance of PD-L1(+) Exosomes in Plasma of Head and Neck Cancer Patients. Clin Cancer Res 2018;24:896-905. [Crossref] [PubMed]
- Poggio M, Hu T, Pai CC, et al. Suppression of Exosomal PD-L1 Induces Systemic Anti-tumor Immunity and Memory. Cell 2019;177:414-27.e13. [Crossref] [PubMed]
- Ludwig S, Floros T, Theodoraki MN, et al. Suppression of Lymphocyte Functions by Plasma Exosomes Correlates with Disease Activity in Patients with Head and Neck Cancer. Clin Cancer Res 2017;23:4843-54. [Crossref] [PubMed]
- Rissin DM, Fournier DR, Piech T, et al. Simultaneous detection of single molecules and singulated ensembles of molecules enables immunoassays with broad dynamic range. Anal Chem 2011;83:2279-85. [Crossref] [PubMed]
- Wilson DH, Rissin DM, Kan CW, et al. The Simoa HD-1 Analyzer: A Novel Fully Automated Digital Immunoassay Analyzer with Single-Molecule Sensitivity and Multiplexing. J Lab Autom 2016;21:533-47. [Crossref] [PubMed]
- Chang L, Rissin DM, Fournier DR, et al. Single molecule enzyme-linked immunosorbent assays: theoretical considerations. J Immunol Methods 2012;378:102-15. [Crossref] [PubMed]
- Rivnak AJ, Rissin DM, Kan CW, et al. A fully-automated, six-plex single molecule immunoassay for measuring cytokines in blood. J Immunol Methods 2015;424:20-7. [Crossref] [PubMed]
- Wei P, Wu F, Kang B, et al. Plasma extracellular vesicles detected by Single Molecule array technology as a liquid biopsy for colorectal cancer. J Extracell Vesicles 2020;9:1809765 [Crossref] [PubMed]
- Baeuerle PA, Gires O. EpCAM (CD326) finding its role in cancer. Br J Cancer 2007;96:417-23. [Crossref] [PubMed]
- Lu SH, Tsai WS, Chang YH, et al. Identifying cancer origin using circulating tumor cells. Cancer Biol Ther 2016;17:430-8. [Crossref] [PubMed]
- Van Der Kraak L, Goel G, Ramanan K, et al. 5-Fluorouracil upregulates cell surface B7-H1 (PD-L1) expression in gastrointestinal cancers. J Immunother Cancer 2016;4:65. [Crossref] [PubMed]
- Kowal J, Arras G, Colombo M, et al. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proc Natl Acad Sci U S A 2016;113:E968-77. [Crossref] [PubMed]
- Bausero MA, Gastpar R, Multhoff G, et al. Alternative mechanism by which IFN-gamma enhances tumor recognition: active release of heat shock protein 72. J Immunol 2005;175:2900-12. [Crossref] [PubMed]
- Theodoraki MN, Hong CS, Donnenberg VS, et al. Evaluation of Exosome Proteins by on-Bead Flow Cytometry. Cytometry A 2021;99:372-81. [Crossref] [PubMed]
- Zheng Q, Huang Y, Zeng X, et al. Correlation between PD-L1 Expression and Clinicopathological and Molecular Characteristics of Non-Small Cell Lung Cancer: A Large Scale Multi-Centric Real-World Study of Chinese Cohort. Laboratory Investigation 2020;100:1834-5.
- Zhu S, Ma L, Wang S, et al. Light-scattering detection below the level of single fluorescent molecules for high-resolution characterization of functional nanoparticles. ACS Nano 2014;8:10998-1006. [Crossref] [PubMed]


